FTDNA versus 23andMe: Shared DNA
This shared DNA is broken down into 3 categories.
1) Sum shared DNA
The sum of shared DNA is the total number of cM (centimorgans) you and a match share across your entire autosomal genome. The total number of cM is approximately 3380cM. Sum shared DNA is typically used for determining close and immediate relationships.
- A parent-child match will share around 3380cM. This is slightly misleading because of course a parent and a child only share 50% of their DNA - the other 50% comes from the other parent. But since this test does not differentiate both pairs of each chromosome, a parent and child will share a SNP at every marker. This is called a half-match.
- Identical twins will also share 3380cM according to the FF results, but since they share 100% of their DNA, those 3380cM will be a full-match - that is, both SNPs at each marker will match compared to only a single SNP between a parent and child.
- Now full-siblings, even though they share 50% of their DNA (same as a parent-child), because each child may have inherited opposite SNPs from each parent at any given marker, full-siblings will generally share less than 3380cM but no less than 2500cM.
- Half-siblings share 25% of their DNA, but like full-siblings it's not going to be exact because the siblings may have inherited opposite SNPs from their shared parent. But the shared DNA for half-siblings should be around 1690cM, and probably no less than 1000cM.
According to Matt Dexter, on the FTDNA Forum, here is a breakdown of sum shared DNA ranges for various relationships:
- Parent/child: 3539-3748 centimorgans (cMs)
- 1st cousins: 548-1034 cMs
- 1st cousins once removed: 248-638 cMs
- 2nd cousins: 101-378 cMs
- 2nd cousins once removed: 43-191 cMs
- 3rd cousins: 43-ca 150 cMs
- 3rd cousins once removed: 11.5-99 cMs
- 4th and more distant cousins: 5-ca 50 cMs
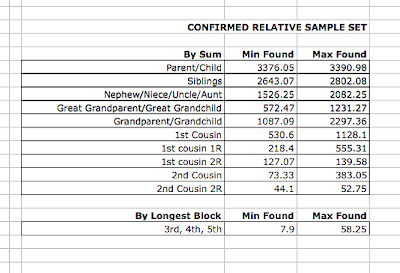
He also provided this great spreadsheet of min and max numbers:
Lastly, it must be noted that usually anything less than 5cM is considered noise. Between 5-10cM suggested possible recent shared ancestry. Over 10cM strongly suggests recent shared ancestry.
2) Longest block of DNA
The longest block of DNA is another tool used primarily for determining more distant relationships.
An explanation of the differences between sum and longest block of DNA from Matt Dexter (FTDNA Forum):
The amount shared is less important for a 4th cousin than for a 1st cousin. The amount shared in a first cousin is a large quantity so it can be used to make a prediction. The amount shared in a 4th cousin is a smaller quantity so it is easier for the test to use the longest block to make a prediction from and then use the sum as a kind of secondary, fine tuning adjustment.And from Tim Janzen on the Genealogy-DNA listserv (http://archiver.rootsweb.
Immediate family have longest blocks however they can't be used because they are different sizes per chromosome. In other words a match on chromosome 1 might override a bigger match on chromosome 3 because chromosome 1 is larger in cM. A parent and child will have the longest block on chromosome 1 for just about 145.17cM but the fact is, all the other chromosomes match too so using the longest block is meaningless for parents, uncles, aunts, nieces, nephews, grandchildren and grandparents for example.
Ranges of the length of shared IBD segments based on family relationship:
- Parent/child: 2851st cousins: 50-141
- 1st cousins once removed: 36-106 cMs
- 2nd cousins: 21-64 cMs
- 2nd cousins once removed: 19-81
- 3rd cousins: 13-77
- 3rd cousins once removed: 0-27
- 4th cousins: 0-22
- 4th cousins once removed: 0-13
- 5th cousins: 0-27
3) Number of segments
The number of segments is not very important, as of course large segments take up more space so immediate relatives will share less numbers of segments but longer segments. The number of segments feature is more useful when looking at more distant relationships, especially where there might only be one significant block of DNA shared.
B. 23andMe's Relative Finder uses shared DNA as well. However, their results are slightly different from FTDNA. First off, only those who have set their profile to public will you see their names (I've discussed this previously). Second, if you have an immediate relative match (parent-child, sibling, grandparent-grandchild, niece/nephew-aunt/uncle, and possibly 1st cousin - I'm not 100% sure on the 1st cousin), those matches will NOT SHOW UP unless that match has signed off to be visible to immediate matches. Why this is, I have no clue. But it's something I was informed of recently from other DC adults who have tested with 23andMe.
Shared DNA from 23andMe is listed in percentages. Which for most people is more understandable, but unclear if you are trying to compare the two companies. Since percentages for two related individuals are fairly contant, it needs little extra information.
DNA Percentages (http://www.isogg.org/wiki/Autosomal_DNA_statistics)
The following figures show the average amount of autosomal DNA shared with close relatives:
- 50% mother, father and siblings
- 25% grandfathers, grandmothers, aunts, uncles, half-siblings, double first cousins
- 12.5% first cousins
- 6.25% first cousins once removed
- 3.125% second cousins, first cousins twice removed
- 0.781% third cousins
- 0.195% fourth cousins
- 0.0488% fifth cousins
- 0.0122% sixth cousins
- 0.00305% seventh cousins (ca 92,000 base pairs)
- 0.000763% eighth cousins (ca 23,000 base pairs)
Of course these numbers can fluctuate slightly due to the randomness of inheritance.
Here are ranges of percentages of genome in common from 23andMe:
- Parent/child: 47.54 (for father/son pairs, who do not share the X chromosome) to ~50%
- 1st cousins: 7.31-13.8
- 1st cousins once removed: 3.3-8.51
- 2nd cousins: 2.85-5.04
- 2nd cousins once removed: .57-2.54
- 3rd cousins: ca .3-2.0
- 3rd cousins once removed: .11-1.32
- 4th and more distant cousins: .07-.5
And just for comparison, here's a breakdown of just how much DNA is shared between UNRELATED people:
- Parent-child pairs share between 83.94% and 84.20% of SNPs (50% of DNA in common)
- Siblings share between 83.81% and 87.47% of SNPs (50% of DNA in common)
- Uncle/aunt-niece/nephew pairs share between 78.48% and 79.57% of SNPs (25% of DNA in common)
- Grandparent-grandchild pairs share between 77.96% and 80.59% of SNPs (12.5% of DNA in common)
- First cousins and great uncle/great aunt-grandniece/grandnephew pairs share 75.78% and 77.03% of SNPs (12.5% of DNA in common)
- First cousins once removed share ca 75.5% of SNPs (6.25% of DNA in common)
- Second cousins and first cousins twice removed share ca 75% of SNPs (3.125% of DNA in common)
- Unrelated people of European descent share 73-74.6% of SNPs

3 comments:
Thank you for your information!
Thanks for this extremely useful information - very much appreciated!
I have a question: is there any reliable way of converting centiMorgans to "percent of shared DNA"? I would have thought no after reading at the top of your article about the inability of the Family Finder test to differentiate between half- and full-matches. But then I was surprised to read further down that Relative Finder reports shared DNA in percentages!? How do they do this?
Thanks
Maurice
I could have just said that I like docmo18's comment, since there is no choice, I had to say it as a comment. Thank you, very much.
Post a Comment